Aminoallyl-dUTP-5/6-TAMRA
Tetramethyl-Rhodamine-5-dUTP
5-(3-Aminoallyl)-2'-deoxyuridine-5'triphosphate, labeled with 5/6-TAMRA, Triethylammonium salt
| Catálogo Nº | Apresentação | Preço (R$) | Comprar / Observação |
|---|---|---|---|
| NU-803-TAM | 30 μl (1 mM) | Sob demanda | Adicionar ao Carrinho |

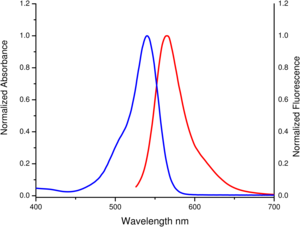
excitation and emission spectrum of 5/6-TAMRA
For general laboratory use.
Envio: shipped on gel packs
Condições de armazenamento: store at -20 °C
Short term exposure (up to 1 week cumulative) to ambient temperature possible.
Validade: 12 months after date of delivery
Fórmula molecular: C37H40N5O18P3 (free acid)
Peso molecular: 935.66 g/mol (free acid)
Pureza: ≥ 95 % (HPLC)
Forma: filtered solution (30 kDa) in 10 mM Tris-HCl
Concentração: 1.0 mM - 1.1 mM
pH: 7.5 ±0.5
Propriedades espectroscópicas: λabs 545 nm, λem 575 nm, ε 90.0 L mmol-1 cm-1 (Tris-HCl pH 7.5)
Formulários:
Incorporation into DNA/cDNA by
- Nick Translation with DNAse I/ DNA Polymerase I in-house data, [1,2]
Descrição:
Aminoallyl-dUTP-5/6-TAMRA is recommended for direct enzymatic labeling of DNA/cDNA by Nick Translation. It is incorporated as substitute for its natural counterpart dTTP. The resulting Dye-labeled DNA/cDNA probes are ideally suited for fluorescence hybridization applications such as FISH or microarray-based gene expression profiling. Optimal substrate properties and thus labeling efficiency is ensured by an optimized linker attached to the C5 position of uridine.
Recommended Aminoallyl-dUTP-5/6-TAMRA/dTTP ratio for Nick Translation: 35% Aminoallyl-dUTP-5/6-TAMRA/ 65% dTTP
Please note: Protect the Dye-labeled dUTP from exposure to light and carry out experimental procedures in low light conditions. The optimal final concentration of the Dye-labeled dUTP may very depending on the application and assay conditions. For optimal produdct yields and high incorporation rates an individual optimization of the Dye-labeled-dUTP/dTTP ratio is recommended.
Referências selecionadas:
[1] Idziak et al. (2014) Insight into the Karyotype Evolution of Brachypodium Species Using Comparative Chromosome Barcoding. PLOS One 9(3):e93503.
[2] Hasterok et al. (2006) Alignment of the Genomes of Brachypodium distachyon and Temperate Cereals and Grasses Using Bacterial Artificial Chromosome Landing With Fluorescence in Situ Hybridization. Genetics 173:349.
